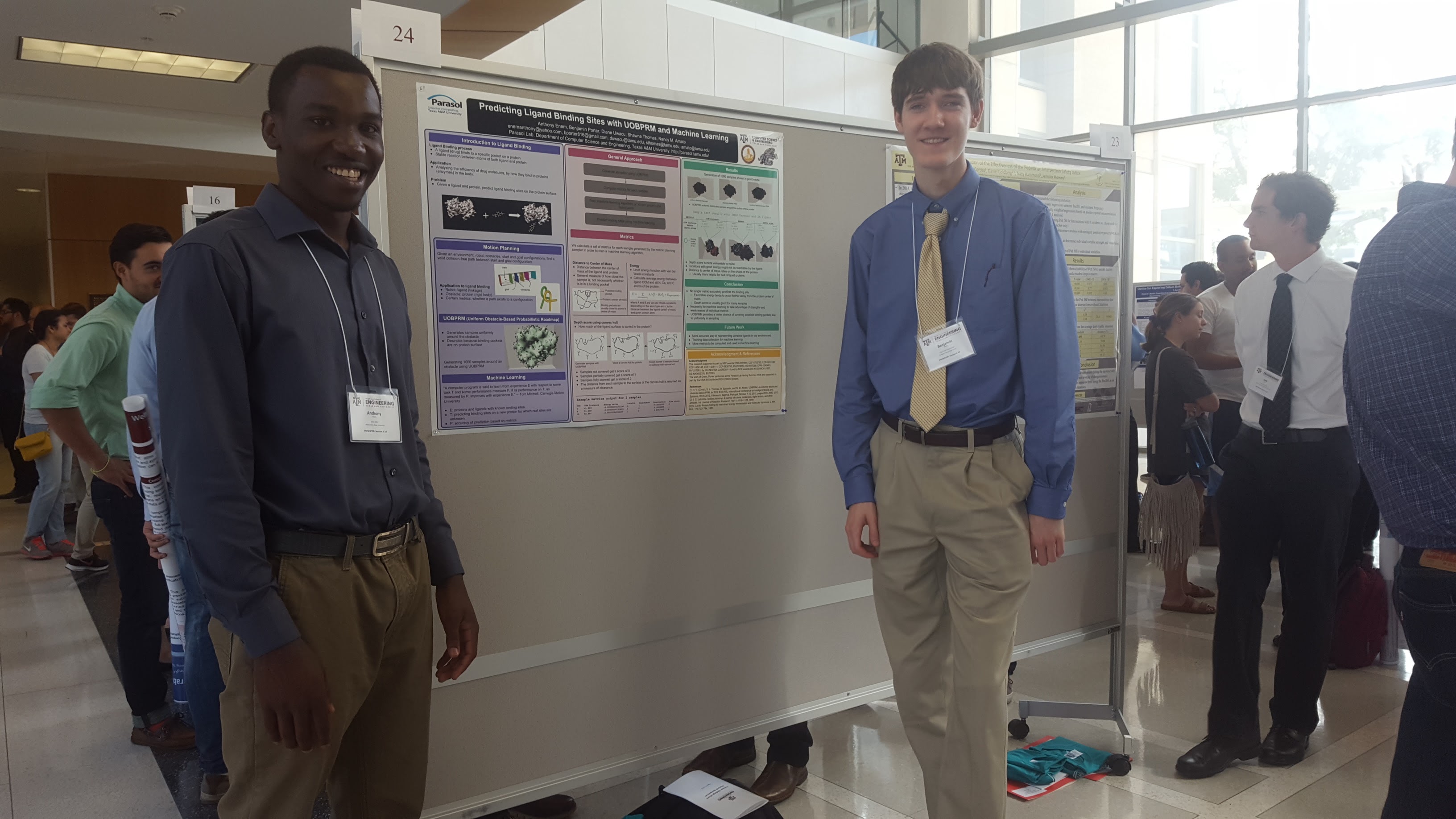
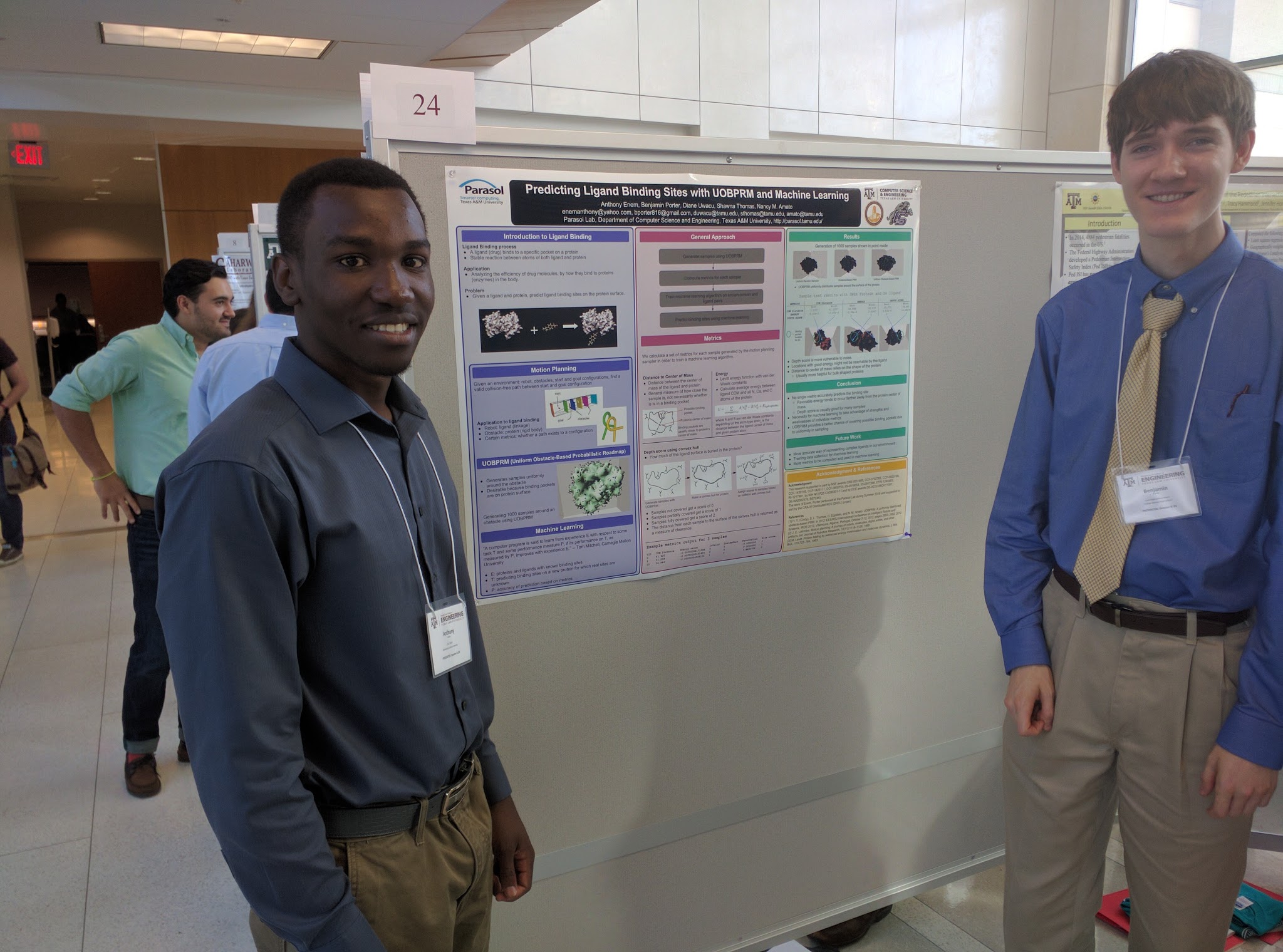
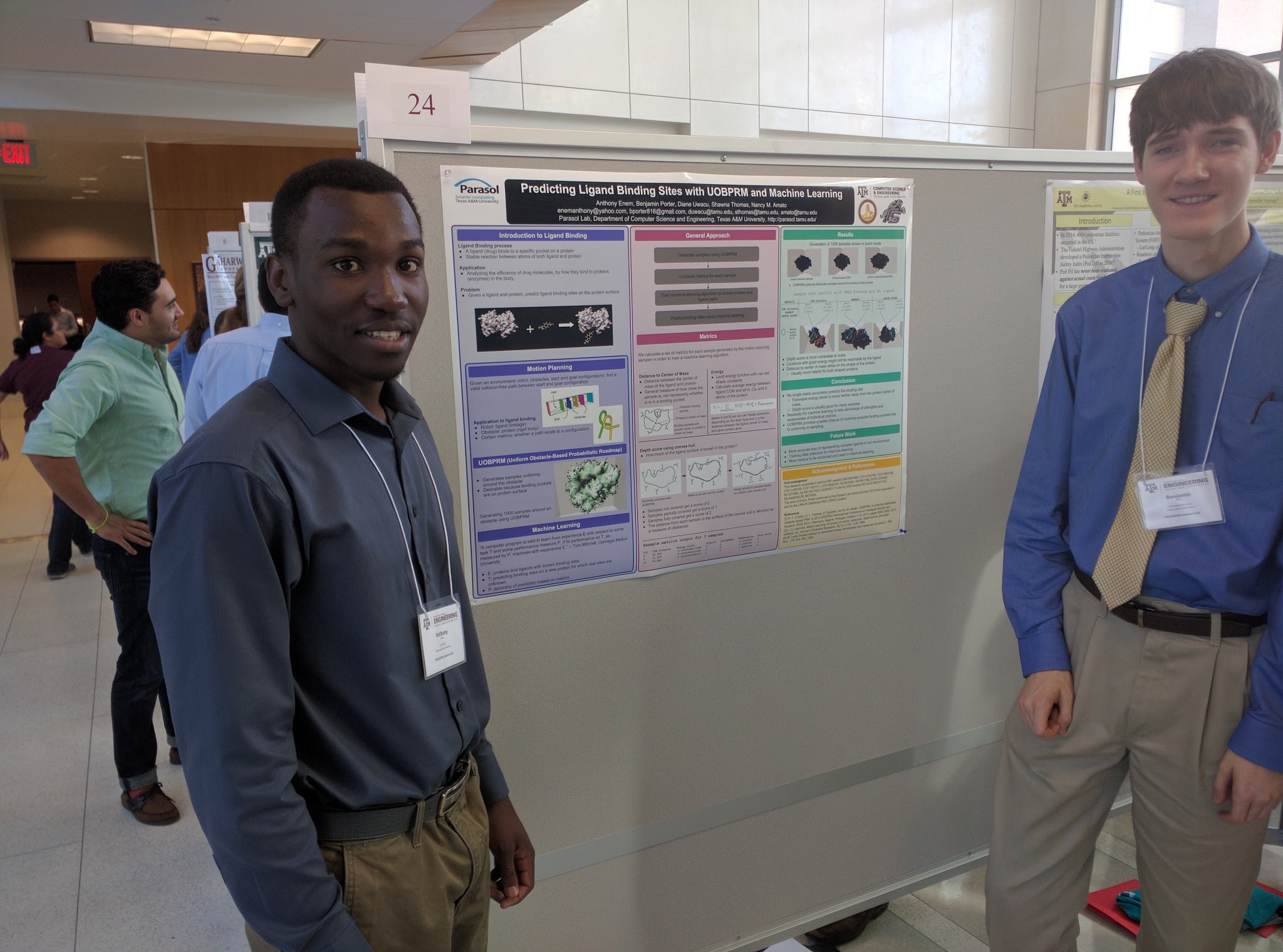
Many approaches to predicting ligand binding sites on protein surfaces suffer from a reliance on unreliable and inaccurate evaluations of potential binding sites. In this work, we propose an approach to ligand binding that takes advantage of the motion planning idea of randomized sampling to uniformly generate samples of the ligand at various binding site candidates near the protein surface. We then compute metrics that describe the favorability of each configuration's location as a potential binding site. We also propose future work to apply these metrics to a machine learning algorithm to take advantage of each metric's strengths and weaknesses to predict the locations of binding sites for specific ligands on the surface of a protein.
Click here to go back to my weekly reports
Click here to go back to my homepage